| 1 |
Loss of function mutations in VARS encoding cytoplasmic valyl-tRNA synthetase cause microcephaly, seizures, and progressive cerebral atrophy.Hum Genet. 2018 Apr;137(4):293-303. doi: 10.1007/s00439-018-1882-3. Epub 2018 Apr 24.
|
| 2 |
URL: http://www.guidetopharmacology.org Nucleic Acids Res. 2015 Oct 12. pii: gkv1037. The IUPHAR/BPS Guide to PHARMACOLOGY in 2016: towards curated quantitative interactions between 1300 protein targets and 6000 ligands. (Ligand id: 5692).
|
| 3 |
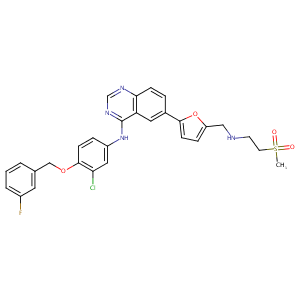
Lapatinib FDA Label
|
| 4 |
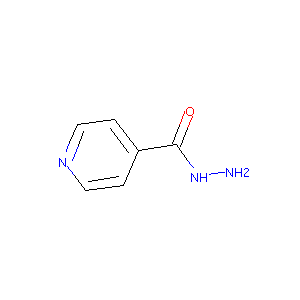
Isoniazid FDA Label
|
| 5 |
Novel agents in the management of Mycobacterium tuberculosis disease. Curr Med Chem. 2007;14(18):2000-8.
|
| 6 |
UGT-dependent regioselective glucuronidation of ursodeoxycholic acid and obeticholic acid and selective transport of the consequent acyl glucuronides by OATP1B1 and 1B3. Chem Biol Interact. 2019 Sep 1;310:108745. doi: 10.1016/j.cbi.2019.108745. Epub 2019 Jul 9.
|
| 7 |
Triple negative breast cancer--current status and prospective targeted treatment based on HER1 (EGFR), TOP2A and C-MYC gene assessment. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub. 2009 Mar;153(1):13-7.
|
| 8 |
Inhibition of eEF-2 kinase sensitizes human nasopharyngeal carcinoma cells to lapatinib-induced apoptosis through the Src and Erk pathways.BMC Cancer. 2016 Oct 19;16(1):813.
|
| 9 |
Tarascon Pocket Pharmacopoeia 2018 Classic Shirt-Pocket Edition.
|
| 10 |
The role of efflux and uptake transporters in [N-{3-chloro-4-[(3-fluorobenzyl)oxy]phenyl}-6-[5-({[2-(methylsulfonyl)ethyl]amino}methyl)-2-furyl]-4-quinazolinamine (GW572016, lapatinib) disposition and drug interactions. Drug Metab Dispos. 2008 Apr;36(4):695-701.
|
| 11 |
Mechanism-based inactivation of cytochrome P450 3A4 by lapatinib. Mol Pharmacol. 2010 Oct;78(4):693-703.
|
| 12 |
Clinical pharmacokinetics of tyrosine kinase inhibitors. Cancer Treat Rev. 2009 Dec;35(8):692-706.
|
| 13 |
The dual ErbB1/ErbB2 inhibitor, lapatinib (GW572016), cooperates with tamoxifen to inhibit both cell proliferation- and estrogen-dependent gene expression in antiestrogen-resistant breast cancer. Cancer Res. 2005 Jan 1;65(1):18-25.
|
| 14 |
Interference with bile salt export pump function is a susceptibility factor for human liver injury in drug development. Toxicol Sci. 2010 Dec; 118(2):485-500.
|
| 15 |
P450 3A-catalyzed O-dealkylation of lapatinib induces mitochondrial stress and activates Nrf2. Chem Res Toxicol. 2016 May 16;29(5):784-96.
|
| 16 |
Combining lapatinib (GW572016), a small molecule inhibitor of ErbB1 and ErbB2 tyrosine kinases, with therapeutic anti-ErbB2 antibodies enhances apoptosis of ErbB2-overexpressing breast cancer cells. Oncogene. 2005 Sep 15;24(41):6213-21. doi: 10.1038/sj.onc.1208774.
|
| 17 |
CDK4/6 inhibition provides a potent adjunct to Her2-targeted therapies in preclinical breast cancer models. Genes Cancer. 2014 Jul;5(7-8):261-72. doi: 10.18632/genesandcancer.24.
|
| 18 |
Effects of lapatinib on cell proliferation and apoptosis in NB4 cells. Oncol Lett. 2018 Jan;15(1):235-242. doi: 10.3892/ol.2017.7342. Epub 2017 Nov 3.
|
| 19 |
The involvement of hepatic cytochrome P450s in the cytotoxicity of lapatinib. Toxicol Sci. 2023 Dec 21;197(1):69-78. doi: 10.1093/toxsci/kfad099.
|
| 20 |
Evaluating the Role of Multidrug Resistance Protein 3 (MDR3) Inhibition in Predicting Drug-Induced Liver Injury Using 125 Pharmaceuticals. Chem Res Toxicol. 2017 May 15;30(5):1219-1229. doi: 10.1021/acs.chemrestox.7b00048. Epub 2017 May 4.
|
| 21 |
Suppression of HER2/HER3-mediated growth of breast cancer cells with combinations of GDC-0941 PI3K inhibitor, trastuzumab, and pertuzumab. Clin Cancer Res. 2009 Jun 15;15(12):4147-56. doi: 10.1158/1078-0432.CCR-08-2814. Epub 2009 Jun 9.
|
| 22 |
Cytotoxicity of 34 FDA approved small-molecule kinase inhibitors in primary rat and human hepatocytes. Toxicol Lett. 2018 Jul;291:138-148. doi: 10.1016/j.toxlet.2018.04.010. Epub 2018 Apr 12.
|
| 23 |
Association of CYP1A1 and CYP1B1 inhibition in in vitro assays with drug-induced liver injury. J Toxicol Sci. 2021;46(4):167-176. doi: 10.2131/jts.46.167.
|
| 24 |
The K-Ras effector p38 MAPK confers intrinsic resistance to tyrosine kinase inhibitors by stimulating EGFR transcription and EGFR dephosphorylation. J Biol Chem. 2017 Sep 8;292(36):15070-15079. doi: 10.1074/jbc.M117.779488. Epub 2017 Jul 24.
|
| 25 |
HLA-DQA1*02:01 is a major risk factor for lapatinib-induced hepatotoxicity in women with advanced breast cancer. J Clin Oncol. 2011 Feb 20;29(6):667-73. doi: 10.1200/JCO.2010.31.3197. Epub 2011 Jan 18.
|
| 26 |
Niclosamide inhibits epithelial-mesenchymal transition and tumor growth in lapatinib-resistant human epidermal growth factor receptor 2-positive breast cancer. Int J Biochem Cell Biol. 2016 Feb;71:12-23. doi: 10.1016/j.biocel.2015.11.014. Epub 2015 Nov 28.
|
| 27 |
Characterization of drug-specific signaling between primary human hepatocytes and immune cells. Toxicol Sci. 2017 Jul 1;158(1):76-89.
|
| 28 |
Diversity in enoyl-acyl carrier protein reductases. Cell Mol Life Sci. 2009 May;66(9):1507-17.
|
| 29 |
Inhibition of CYP2E1 catalytic activity in vitro by S-adenosyl-L-methionine. Biochem Pharmacol. 2005 Apr 1;69(7):1081-93.
|
| 30 |
Crystal structure of the catalase-peroxidase KatG W78F mutant from Synechococcus elongatus PCC7942 in complex with the antitubercular pro-drug isoniazid. FEBS Lett. 2015 Jan 2;589(1):131-7.
|
| 31 |
The actinobacterium Tsukamurella paurometabola has a functionally divergent arylamine N-acetyltransferase (NAT) homolog. World J Microbiol Biotechnol. 2019 Oct 31;35(11):174.
|
| 32 |
Quercetin protected against isoniazide-induced HepG2 cell apoptosis by activating the SIRT1/ERK pathway. J Biochem Mol Toxicol. 2019 Sep;33(9):e22369. doi: 10.1002/jbt.22369. Epub 2019 Jul 23.
|
| 33 |
ADReCS-Target: target profiles for aiding drug safety research and application. Nucleic Acids Res. 2018 Jan 4;46(D1):D911-D917. doi: 10.1093/nar/gkx899.
|
| 34 |
Mechanism-based inactivation of human cytochrome P4502C8 by drugs in vitro. J Pharmacol Exp Ther. 2004 Dec;311(3):996-1007.
|
| 35 |
Determination of phospholipidosis potential based on gene expression analysis in HepG2 cells. Toxicol Sci. 2007 Mar;96(1):101-14.
|
| 36 |
Comparison of base-line and chemical-induced transcriptomic responses in HepaRG and RPTEC/TERT1 cells using TempO-Seq. Arch Toxicol. 2018 Aug;92(8):2517-2531.
|
| 37 |
Identification of differentially expressed genes in hepatic HepG2 cells treated with acetaminophen using suppression subtractive hybridization. Biol Pharm Bull. 2005 Jul;28(7):1148-53. doi: 10.1248/bpb.28.1148.
|
| 38 |
Effect of common medications on the expression of SARS-CoV-2 entry receptors in liver tissue. Arch Toxicol. 2020 Dec;94(12):4037-4041. doi: 10.1007/s00204-020-02869-1. Epub 2020 Aug 17.
|
| 39 |
An in vitro coculture system of human peripheral blood mononuclear cells with hepatocellular carcinoma-derived cells for predicting drug-induced liver injury. Arch Toxicol. 2021 Jan;95(1):149-168. doi: 10.1007/s00204-020-02882-4. Epub 2020 Aug 20.
|
| 40 |
Auto-oxidation of Isoniazid Leads to Isonicotinic-Lysine Adducts on Human Serum Albumin. Chem Res Toxicol. 2015 Jan 20;28(1):51-8. doi: 10.1021/tx500285k. Epub 2014 Dec 9.
|
| 41 |
Isoniazid-induced apoptosis in HepG2 cells: generation of oxidative stress and Bcl-2 down-regulation. Toxicol Mech Methods. 2010 Jun;20(5):242-51. doi: 10.3109/15376511003793325.
|
| 42 |
The Isoniazid Metabolites Hydrazine and Pyridoxal Isonicotinoyl Hydrazone Modulate Heme Biosynthesis. Toxicol Sci. 2019 Mar 1;168(1):209-224. doi: 10.1093/toxsci/kfy294.
|
| 43 |
Isoniazid suppresses antioxidant response element activities and impairs adipogenesis in mouse and human preadipocytes. Toxicol Appl Pharmacol. 2013 Dec 15;273(3):435-41. doi: 10.1016/j.taap.2013.10.005. Epub 2013 Oct 12.
|
| 44 |
AMPK activator acadesine fails to alleviate isoniazid-caused mitochondrial instability in HepG2 cells. J Appl Toxicol. 2017 Oct;37(10):1219-1224. doi: 10.1002/jat.3483. Epub 2017 May 29.
|
| 45 |
Enhanced activation of human NK cells by drug-exposed hepatocytes. Arch Toxicol. 2020 Feb;94(2):439-448. doi: 10.1007/s00204-020-02668-8. Epub 2020 Feb 14.
|
| 46 |
Effects of N-acetyltransferase 2 (NAT2), CYP2E1 and Glutathione-S-transferase (GST) genotypes on the serum concentrations of isoniazid and metabolites in tuberculosis patients. J Toxicol Sci. 2008 May;33(2):187-95. doi: 10.2131/jts.33.187.
|
| 47 |
Eosinophil peroxidase oxidizes isoniazid to form the active metabolite against M. tuberculosis, isoniazid-NAD(). Chem Biol Interact. 2019 May 25;305:48-53. doi: 10.1016/j.cbi.2019.03.019. Epub 2019 Mar 25.
|
| 48 |
Metabolism of isoniazid by neutrophil myeloperoxidase leads to isoniazid-NAD(+) adduct formation: A comparison of the reactivity of isoniazid with its known human metabolites. Biochem Pharmacol. 2016 Apr 15;106:46-55. doi: 10.1016/j.bcp.2016.02.003. Epub 2016 Feb 9.
|
| 49 |
Development of a highly sensitive cytotoxicity assay system for CYP3A4-mediated metabolic activation. Drug Metab Dispos. 2011 Aug;39(8):1388-95. doi: 10.1124/dmd.110.037077. Epub 2011 May 3.
|
| 50 |
Customised in vitro model to detect human metabolism-dependent idiosyncratic drug-induced liver injury. Arch Toxicol. 2018 Jan;92(1):383-399. doi: 10.1007/s00204-017-2036-4. Epub 2017 Jul 31.
|
| 51 |
Biologically active neutrophil chemokine pattern in tonsillitis.Clin Exp Immunol. 2004 Mar;135(3):511-8. doi: 10.1111/j.1365-2249.2003.02390.x.
|
|
|
|
|
|
|