| 1 |
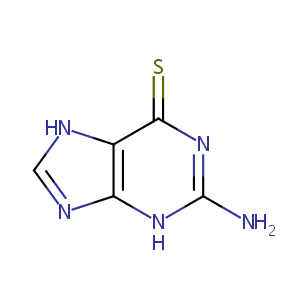
Recurrent recessive mutation in deoxyguanosine kinase causes idiopathic noncirrhotic portal hypertension.Hepatology. 2016 Jun;63(6):1977-86. doi: 10.1002/hep.28499. Epub 2016 Mar 31.
|
| 2 |
Drugs@FDA. U.S. Food and Drug Administration. U.S. Department of Health & Human Services. 2015
|
| 3 |
Thioguanine FDA Label
|
| 4 |
URL: http://www.guidetopharmacology.org Nucleic Acids Res. 2015 Oct 12. pii: gkv1037. The IUPHAR/BPS Guide to PHARMACOLOGY in 2016: towards curated quantitative interactions between 1300 protein targets and 6000 ligands. (Ligand id: 6845).
|
| 5 |
Thiopurine analogs and mycophenolic acid synergistically inhibit the papain-like protease of Middle East respiratory syndrome coronavirus. Antiviral Res. 2015 Mar;115:9-16.
|
| 6 |
URL: http://www.guidetopharmacology.org Nucleic Acids Res. 2015 Oct 12. pii: gkv1037. The IUPHAR/BPS Guide to PHARMACOLOGY in 2016: towards curated quantitative interactions between 1300 protein targets and 6000 ligands. (Ligand id: 5697).
|
| 7 |
Immune effector cells produce lethal DNA damage in cells treated with a thiopurine. Cancer Res. 2009 Mar 15;69(6):2393-9.
|
| 8 |
ABC transporters and their role in nucleoside and nucleotide drug resistance. Biochem Pharmacol. 2012 Apr 15;83(8):1073-83.
|
| 9 |
Multidrug resistance protein 4 (MRP4/ABCC4) mediates efflux of bimane-glutathione. Int J Biochem Cell Biol. 2004 Feb;36(2):247-57.
|
| 10 |
Involvement of the concentrative nucleoside transporter 3 and equilibrative nucleoside transporter 2 in the resistance of T-lymphoblastic cell lines to thiopurines. Biochem Biophys Res Commun. 2006 Apr 28;343(1):208-15.
|
| 11 |
Usefulness of thiopurine methyltransferase and thiopurine metabolite analysis in clinical practice in patients with inflammatory bowel diseases. Acta Gastroenterol Belg. 2010 Jul-Sep;73(3):331-5.
|
| 12 |
Qualitative and quantitative difference in mutation induction between carbon- and neon-ion beams in normal human cells. Biol Sci Space. 2003 Dec;17(4):302-6.
|
| 13 |
ADReCS-Target: target profiles for aiding drug safety research and application. Nucleic Acids Res. 2018 Jan 4;46(D1):D911-D917. doi: 10.1093/nar/gkx899.
|
| 14 |
CRISPR/Cas9-mediated gene knockout screens and target identification via whole-genome sequencing uncover host genes required for picornavirus infection. J Biol Chem. 2017 Jun 23;292(25):10664-10671. doi: 10.1074/jbc.M117.782425. Epub 2017 Apr 26.
|
| 15 |
Clinical Pharmacogenetics Implementation Consortium guidelines for thiopurine methyltransferase genotype and thiopurine dosing. Clin Pharmacol Ther. 2011 Mar;89(3):387-91. doi: 10.1038/clpt.2010.320. Epub 2011 Jan 26.
|
| 16 |
Petit E, Langouet S, Akhdar H, Nicolas-Nicolaz C, Guillouzo A, Morel F. Differential toxic effects of azathioprine, 6-mercaptopurine and 6-thioguanine on human hepatocytes. Toxicol In Vitro. 2008;22(3):632-642. [PMID: 18222062]
|
| 17 |
Important role of caspase-8 for chemosensitivity of ALL cells. Clin Cancer Res. 2011 Dec 15;17(24):7605-13. doi: 10.1158/1078-0432.CCR-11-0513. Epub 2011 Oct 18.
|
| 18 |
The effect of 6-thioguanine on alternative splicing and antisense-mediated exon skipping treatment for duchenne muscular dystrophy. PLoS Curr. 2012 Dec 12;4:ecurrents.md.597d700f92eaa70de261ea0d91821377. doi: 10.1371/currents.md.597d700f92eaa70de261ea0d91821377.
|
| 19 |
The thiopurine methyltransferase genetic polymorphism is associated with thioguanine-related veno-occlusive disease of the liver in children with acute lymphoblastic leukemia. Clin Pharmacol Ther. 2006 Oct;80(4):375-83. doi: 10.1016/j.clpt.2006.07.002.
|
| 20 |
The effect of Msh2 knockdown on methylating agent induced toxicity in DNA glycosylase deficient cells. Toxicology. 2010 Jan 31;268(1-2):111-7. doi: 10.1016/j.tox.2009.12.008. Epub 2009 Dec 16.
|
| 21 |
The multidrug resistance protein 5 (ABCC5) confers resistance to 5-fluorouracil and transports its monophosphorylated metabolites. Mol Cancer Ther. 2005 May;4(5):855-63.
|
| 22 |
Expression and prognostic significance of IAP-family genes in human cancers and myeloid leukemias. Clin Cancer Res. 2000 May;6(5):1796-803.
|
| 23 |
Assessment of the inhibition potential of Licochalcone A against human UDP-glucuronosyltransferases. Food Chem Toxicol. 2016 Apr;90:112-22.
|
| 24 |
Exposure-based assessment of chemical teratogenicity using morphogenetic aggregates of human embryonic stem cells. Reprod Toxicol. 2020 Jan;91:74-91. doi: 10.1016/j.reprotox.2019.10.004. Epub 2019 Nov 8.
|
| 25 |
Endoplasmic reticulum stress-mediated apoptosis in imatinib-resistant leukemic K562-r cells triggered by AMN107 combined with arsenic trioxide. Exp Biol Med (Maywood). 2013 Aug 1;238(8):932-42. doi: 10.1177/1535370213492689. Epub 2013 Jul 24.
|
| 26 |
2007 FDA drug approvals: a year of flux. Nat Rev Drug Discov. 2008 Feb;7(2):107-9.
|
| 27 |
Interaction of nilotinib, dasatinib and bosutinib with ABCB1 and ABCG2: implications for altered anti-cancer effects and pharmacological properties. Br J Pharmacol. 2009 Oct;158(4):1153-64.
|
| 28 |
KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017 Jan 4;45(D1):D353-D361. (dg:DG01665)
|
| 29 |
Contribution of OATP1B1 and OATP1B3 to the disposition of sorafenib and sorafenib-glucuronide. Clin Cancer Res. 2013 Mar 15;19(6):1458-66.
|
| 30 |
Drug interactions with the tyrosine kinase inhibitors imatinib, dasatinib, and nilotinib. Blood. 2011 Feb 24;117(8):e75-87.
|
| 31 |
Role of cytochrome P450 2C8 in drug metabolism and interactions. Pharmacol Rev. 2016 Jan;68(1):168-241.
|
| 32 |
Resistance to daunorubicin, imatinib, or nilotinib depends on expression levels of ABCB1 and ABCG2 in human leukemia cells. Chem Biol Interact. 2014 Aug 5;219:203-10. doi: 10.1016/j.cbi.2014.06.009. Epub 2014 Jun 19.
|
| 33 |
Reversal of ABCB1 mediated efflux by imatinib and nilotinib in cells expressing various transporter levels. Chem Biol Interact. 2017 Aug 1;273:171-179. doi: 10.1016/j.cbi.2017.06.012. Epub 2017 Jun 13.
|
| 34 |
Multi-parameter in vitro toxicity testing of crizotinib, sunitinib, erlotinib, and nilotinib in human cardiomyocytes. Toxicol Appl Pharmacol. 2013 Oct 1;272(1):245-55.
|
| 35 |
p53 Gene (NY-CO-13) Levels in Patients with Chronic Myeloid Leukemia: The Role of Imatinib and Nilotinib. Diseases. 2018 Jan 25;6(1):13. doi: 10.3390/diseases6010013.
|
| 36 |
Nilotinib reduced the viability of human ovarian cancer cells via mitochondria-dependent apoptosis, independent of JNK activation. Toxicol In Vitro. 2016 Mar;31:1-11. doi: 10.1016/j.tiv.2015.11.002. Epub 2015 Nov 6.
|
| 37 |
AP24534, a pan-BCR-ABL inhibitor for chronic myeloid leukemia, potently inhibits the T315I mutant and overcomes mutation-based resistance. Cancer Cell. 2009 Nov 6;16(5):401-12. doi: 10.1016/j.ccr.2009.09.028.
|
| 38 |
The CML-related oncoprotein BCR/ABL induces expression of histidine decarboxylase (HDC) and the synthesis of histamine in leukemic cells. Blood. 2006 Nov 15;108(10):3538-47. doi: 10.1182/blood-2005-12-028456. Epub 2006 Jul 18.
|
| 39 |
Cytotoxicity of 34 FDA approved small-molecule kinase inhibitors in primary rat and human hepatocytes. Toxicol Lett. 2018 Jul;291:138-148. doi: 10.1016/j.toxlet.2018.04.010. Epub 2018 Apr 12.
|
| 40 |
Biologically active neutrophil chemokine pattern in tonsillitis.Clin Exp Immunol. 2004 Mar;135(3):511-8. doi: 10.1111/j.1365-2249.2003.02390.x.
|
| 41 |
Loss of function mutations in VARS encoding cytoplasmic valyl-tRNA synthetase cause microcephaly, seizures, and progressive cerebral atrophy.Hum Genet. 2018 Apr;137(4):293-303. doi: 10.1007/s00439-018-1882-3. Epub 2018 Apr 24.
|
|
|
|
|
|
|