| 1 |
Recurrent recessive mutation in deoxyguanosine kinase causes idiopathic noncirrhotic portal hypertension.Hepatology. 2016 Jun;63(6):1977-86. doi: 10.1002/hep.28499. Epub 2016 Mar 31.
|
| 2 |
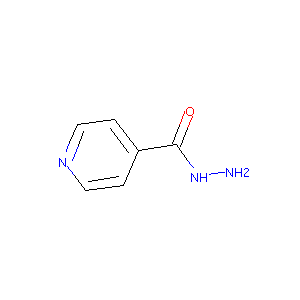
Isoniazid FDA Label
|
| 3 |
Novel agents in the management of Mycobacterium tuberculosis disease. Curr Med Chem. 2007;14(18):2000-8.
|
| 4 |
URL: http://www.guidetopharmacology.org Nucleic Acids Res. 2015 Oct 12. pii: gkv1037. The IUPHAR/BPS Guide to PHARMACOLOGY in 2016: towards curated quantitative interactions between 1300 protein targets and 6000 ligands. (Ligand id: 7143).
|
| 5 |
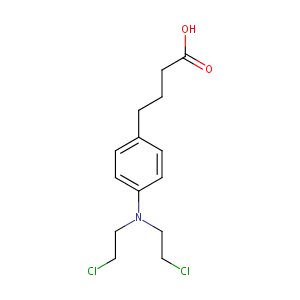
Chlorambucil FDA Label
|
| 6 |
Characterization of drug-specific signaling between primary human hepatocytes and immune cells. Toxicol Sci. 2017 Jul 1;158(1):76-89.
|
| 7 |
Diversity in enoyl-acyl carrier protein reductases. Cell Mol Life Sci. 2009 May;66(9):1507-17.
|
| 8 |
Inhibition of CYP2E1 catalytic activity in vitro by S-adenosyl-L-methionine. Biochem Pharmacol. 2005 Apr 1;69(7):1081-93.
|
| 9 |
Crystal structure of the catalase-peroxidase KatG W78F mutant from Synechococcus elongatus PCC7942 in complex with the antitubercular pro-drug isoniazid. FEBS Lett. 2015 Jan 2;589(1):131-7.
|
| 10 |
The actinobacterium Tsukamurella paurometabola has a functionally divergent arylamine N-acetyltransferase (NAT) homolog. World J Microbiol Biotechnol. 2019 Oct 31;35(11):174.
|
| 11 |
Quercetin protected against isoniazide-induced HepG2 cell apoptosis by activating the SIRT1/ERK pathway. J Biochem Mol Toxicol. 2019 Sep;33(9):e22369. doi: 10.1002/jbt.22369. Epub 2019 Jul 23.
|
| 12 |
ADReCS-Target: target profiles for aiding drug safety research and application. Nucleic Acids Res. 2018 Jan 4;46(D1):D911-D917. doi: 10.1093/nar/gkx899.
|
| 13 |
Mechanism-based inactivation of human cytochrome P4502C8 by drugs in vitro. J Pharmacol Exp Ther. 2004 Dec;311(3):996-1007.
|
| 14 |
Determination of phospholipidosis potential based on gene expression analysis in HepG2 cells. Toxicol Sci. 2007 Mar;96(1):101-14.
|
| 15 |
Comparison of base-line and chemical-induced transcriptomic responses in HepaRG and RPTEC/TERT1 cells using TempO-Seq. Arch Toxicol. 2018 Aug;92(8):2517-2531.
|
| 16 |
Identification of differentially expressed genes in hepatic HepG2 cells treated with acetaminophen using suppression subtractive hybridization. Biol Pharm Bull. 2005 Jul;28(7):1148-53. doi: 10.1248/bpb.28.1148.
|
| 17 |
Effect of common medications on the expression of SARS-CoV-2 entry receptors in liver tissue. Arch Toxicol. 2020 Dec;94(12):4037-4041. doi: 10.1007/s00204-020-02869-1. Epub 2020 Aug 17.
|
| 18 |
An in vitro coculture system of human peripheral blood mononuclear cells with hepatocellular carcinoma-derived cells for predicting drug-induced liver injury. Arch Toxicol. 2021 Jan;95(1):149-168. doi: 10.1007/s00204-020-02882-4. Epub 2020 Aug 20.
|
| 19 |
Auto-oxidation of Isoniazid Leads to Isonicotinic-Lysine Adducts on Human Serum Albumin. Chem Res Toxicol. 2015 Jan 20;28(1):51-8. doi: 10.1021/tx500285k. Epub 2014 Dec 9.
|
| 20 |
Isoniazid-induced apoptosis in HepG2 cells: generation of oxidative stress and Bcl-2 down-regulation. Toxicol Mech Methods. 2010 Jun;20(5):242-51. doi: 10.3109/15376511003793325.
|
| 21 |
The Isoniazid Metabolites Hydrazine and Pyridoxal Isonicotinoyl Hydrazone Modulate Heme Biosynthesis. Toxicol Sci. 2019 Mar 1;168(1):209-224. doi: 10.1093/toxsci/kfy294.
|
| 22 |
Isoniazid suppresses antioxidant response element activities and impairs adipogenesis in mouse and human preadipocytes. Toxicol Appl Pharmacol. 2013 Dec 15;273(3):435-41. doi: 10.1016/j.taap.2013.10.005. Epub 2013 Oct 12.
|
| 23 |
AMPK activator acadesine fails to alleviate isoniazid-caused mitochondrial instability in HepG2 cells. J Appl Toxicol. 2017 Oct;37(10):1219-1224. doi: 10.1002/jat.3483. Epub 2017 May 29.
|
| 24 |
Enhanced activation of human NK cells by drug-exposed hepatocytes. Arch Toxicol. 2020 Feb;94(2):439-448. doi: 10.1007/s00204-020-02668-8. Epub 2020 Feb 14.
|
| 25 |
Effects of N-acetyltransferase 2 (NAT2), CYP2E1 and Glutathione-S-transferase (GST) genotypes on the serum concentrations of isoniazid and metabolites in tuberculosis patients. J Toxicol Sci. 2008 May;33(2):187-95. doi: 10.2131/jts.33.187.
|
| 26 |
Eosinophil peroxidase oxidizes isoniazid to form the active metabolite against M. tuberculosis, isoniazid-NAD(). Chem Biol Interact. 2019 May 25;305:48-53. doi: 10.1016/j.cbi.2019.03.019. Epub 2019 Mar 25.
|
| 27 |
Metabolism of isoniazid by neutrophil myeloperoxidase leads to isoniazid-NAD(+) adduct formation: A comparison of the reactivity of isoniazid with its known human metabolites. Biochem Pharmacol. 2016 Apr 15;106:46-55. doi: 10.1016/j.bcp.2016.02.003. Epub 2016 Feb 9.
|
| 28 |
Development of a highly sensitive cytotoxicity assay system for CYP3A4-mediated metabolic activation. Drug Metab Dispos. 2011 Aug;39(8):1388-95. doi: 10.1124/dmd.110.037077. Epub 2011 May 3.
|
| 29 |
Customised in vitro model to detect human metabolism-dependent idiosyncratic drug-induced liver injury. Arch Toxicol. 2018 Jan;92(1):383-399. doi: 10.1007/s00204-017-2036-4. Epub 2017 Jul 31.
|
| 30 |
Roles of DNA repair and reductase activity in the cytotoxicity of the hypoxia-activated dinitrobenzamide mustard PR-104A. Mol Cancer Ther. 2009 Jun;8(6):1714-23.
|
| 31 |
Mammalian drug efflux transporters of the ATP binding cassette (ABC) family in multidrug resistance: A review of the past decade. Cancer Lett. 2016 Jan 1;370(1):153-64.
|
| 32 |
Transporters and renal drug elimination. Annu Rev Pharmacol Toxicol. 2004;44:137-66.
|
| 33 |
The anti-cancer drug chlorambucil as a substrate for the human polymorphic enzyme glutathione transferase P1-1: kinetic properties and crystallographic characterisation of allelic variants. J Mol Biol. 2008 Jun 27;380(1):131-44.
|
| 34 |
The influence of coordinate overexpression of glutathione phase II detoxification gene products on drug resistance. J Pharmacol Exp Ther. 2000 Aug;294(2):480-7.
|
| 35 |
Enhanced in vitro invasiveness and drug resistance with altered gene expression patterns in a human lung carcinoma cell line after pulse selection with anticancer drugs. Int J Cancer. 2004 Sep 10;111(4):484-93. doi: 10.1002/ijc.20230.
|
| 36 |
Role of multidrug resistance protein 1 (MRP1) and glutathione S-transferase A1-1 in alkylating agent resistanceKinetics of glutathione conjugate formation and efflux govern differential cellular sensitivity to chlorambucil versus melphalan toxicity. J Biol Chem. 2001 Mar 16;276(11):7952-6.
|
| 37 |
Oxidative stress mechanisms do not discriminate between genotoxic and nongenotoxic liver carcinogens. Chem Res Toxicol. 2015 Aug 17;28(8):1636-46.
|
| 38 |
Role of the TRAIL/APO2-L death receptors in chlorambucil- and fludarabine-induced apoptosis in chronic lymphocytic leukemia. Oncogene. 2003 Nov 13;22(51):8356-69. doi: 10.1038/sj.onc.1207004.
|
| 39 |
Differential effects of chemotherapeutic drugs versus the MDM-2 antagonist nutlin-3 on cell cycle progression and induction of apoptosis in SKW6.4 lymphoblastoid B-cells. J Cell Biochem. 2008 May 15;104(2):595-605. doi: 10.1002/jcb.21649.
|
| 40 |
Differential gene expression induction by TRAIL in B chronic lymphocytic leukemia (B-CLL) cells showing high versus low levels of Zap-70. J Cell Physiol. 2007 Oct;213(1):229-36. doi: 10.1002/jcp.21116.
|
| 41 |
Theophylline synergizes with chlorambucil in inducing apoptosis of B-chronic lymphocytic leukemia cells. Blood. 1996 Sep 15;88(6):2172-82.
|
| 42 |
A two-hit mechanism for pre-mitotic arrest of cancer cell proliferation by a polyamide-alkylator conjugate. Cell Cycle. 2006 Jul;5(14):1537-48. doi: 10.4161/cc.5.14.2913. Epub 2006 Jul 17.
|
| 43 |
Systems pharmacological analysis of drugs inducing stevens-johnson syndrome and toxic epidermal necrolysis. Chem Res Toxicol. 2015 May 18;28(5):927-34. doi: 10.1021/tx5005248. Epub 2015 Apr 3.
|
| 44 |
Caspase 8 activation independent of Fas (CD95/APO-1) signaling may mediate killing of B-chronic lymphocytic leukemia cells by cytotoxic drugs or gamma radiation. Blood. 2001 Nov 1;98(9):2800-7. doi: 10.1182/blood.v98.9.2800.
|
| 45 |
Disruption of gene expression and induction of apoptosis in prostate cancer cells by a DNA-damaging agent tethered to an androgen receptor ligand. Chem Biol. 2005 Jul;12(7):779-87. doi: 10.1016/j.chembiol.2005.05.009.
|
| 46 |
Chlorambucil induction of HsRad51 in B-cell chronic lymphocytic leukemia. Clin Cancer Res. 1999 Aug;5(8):2178-84.
|
| 47 |
Bcl-2 antisense oligonucleotides enhance the cytotoxicity of chlorambucil in B-cell chronic lymphocytic leukaemia cells. Leuk Lymphoma. 2001 Jul;42(3):491-8. doi: 10.3109/10428190109064606.
|
| 48 |
The MT1G Gene in LUHMES Neurons Is a Sensitive Biomarker of Neurotoxicity. Neurotox Res. 2020 Dec;38(4):967-978. doi: 10.1007/s12640-020-00272-3. Epub 2020 Sep 1.
|
| 49 |
Characterization of the cancer chemopreventive NRF2-dependent gene battery in human keratinocytes: demonstration that the KEAP1-NRF2 pathway, and not the BACH1-NRF2 pathway, controls cytoprotection against electrophiles as well as redox-cycling compounds. Carcinogenesis. 2009 Sep;30(9):1571-80. doi: 10.1093/carcin/bgp176. Epub 2009 Jul 16.
|
| 50 |
Increased resistance to nitrogen mustards and antifolates following in vitro selection of murine fibroblasts and primary hematopoietic cells transduced with a bicistronic retroviral vector expressing the rat glutathione S-transferase A3 and a mutant dihydrofolate reductase. Cancer Gene Ther. 2003 Aug;10(8):637-46. doi: 10.1038/sj.cgt.7700619.
|
| 51 |
Coexpression of rat glutathione S-transferase A3 and human cytidine deaminase by a bicistronic retroviral vector confers in vitro resistance to nitrogen mustards and cytosine arabinoside in murine fibroblasts. Cancer Gene Ther. 2000 May;7(5):757-65. doi: 10.1038/sj.cgt.7700169.
|
| 52 |
In vivo therapeutic responses contingent on Fanconi anemia/BRCA2 status of the tumor. Clin Cancer Res. 2005 Oct 15;11(20):7508-15. doi: 10.1158/1078-0432.CCR-05-1048.
|
| 53 |
Expression and prognostic significance of IAP-family genes in human cancers and myeloid leukemias. Clin Cancer Res. 2000 May;6(5):1796-803.
|
| 54 |
Role of multidrug resistance protein 2 (MRP2, ABCC2) in alkylating agent detoxification: MRP2 potentiates glutathione S-transferase A1-1-mediated resistance to chlorambucil cytotoxicity. J Pharmacol Exp Ther. 2004 Jan;308(1):260-7. doi: 10.1124/jpet.103.057729. Epub 2003 Oct 20.
|
| 55 |
Glutathione S-transferase M1 and multidrug resistance protein 1 act in synergy to protect melanoma cells from vincristine effects. Mol Pharmacol. 2004 Apr;65(4):897-905. doi: 10.1124/mol.65.4.897.
|
|
|
|
|
|
|